 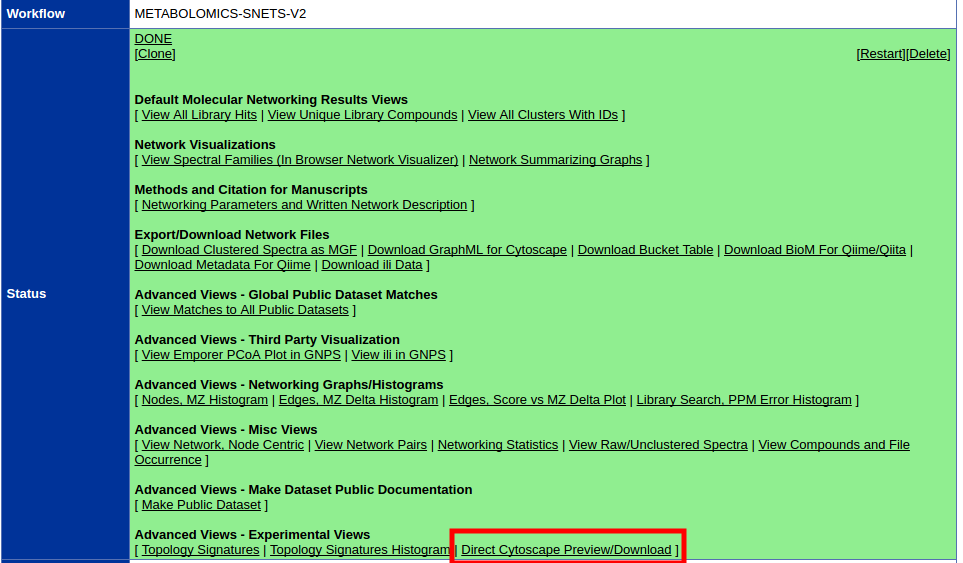 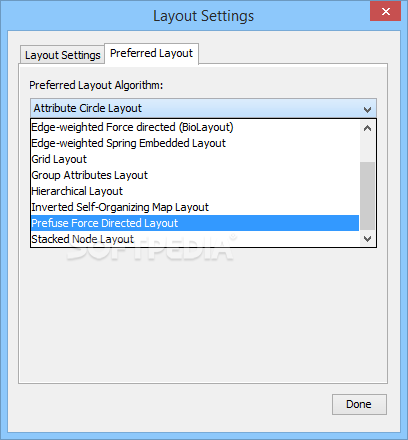
To move the window around the network, you can either use the middle mouseīutton, or drag the small window ���outlined in blue around in the Network Overview Pane.To zoom in on the selected nodes, click on the icon.Again, a group of nodes will be selected and can be moved around on the screen. To select a group of nodes, hold the mouse down in the upper left-hand corner and drag your mouse over a region of the network.Note that both nodes are now selected (yellow). Now add another node to the selection by holding down the Shift key and clicking on a node.If you hold your mouse down over the node and drag it around the node will move on the screen. Start by clicking on the node at the upper left corner of the network.Manipulating Your Network Now that you have a network loaded, you can interact with it in a number of ways: Source and target are gene/protein identifiers that are used to define nodes, while interaction type serves to label the edge connecting each pair of nodes. It consists of 3 columns: source, interaction type, and target. The SIF file format is about as simple as it gets. Open the sampleData folder and select galFiltered.sif and then click on Open and then.For Data Source Type select Local and then click Select.You should see the Import Network File Dialog.Go to File-> Import -> Network (multiple file types).The About option displays information about the running version of Cytoscape.The Help menu allows you to launch the help viewer and browse the table of contents for.Depending on which plugins are loaded, the plugins that you see may be different than what appear here.Plugins The Plugins menu contains options for managing your plugins (install/update/delete) and may have options added by plugins that have already been installed, such as the Agilent Literature Search or Merge Networks. The bottom section of the menu lists a variety of layout algorithms that automatically lay.Rotate, Scale, Align and Distribute are tools for manipulating the network visualization.Layout The Layout menu has an array of features for visually organizing the network: Portions of a network whose node or edge attributes meet a filtering criterion (see below for the filters section). The Select à Use Filters option allows filters to be created for automatic selection of.The network management panel (Control Panel).View The View menu allows you to display or hide: 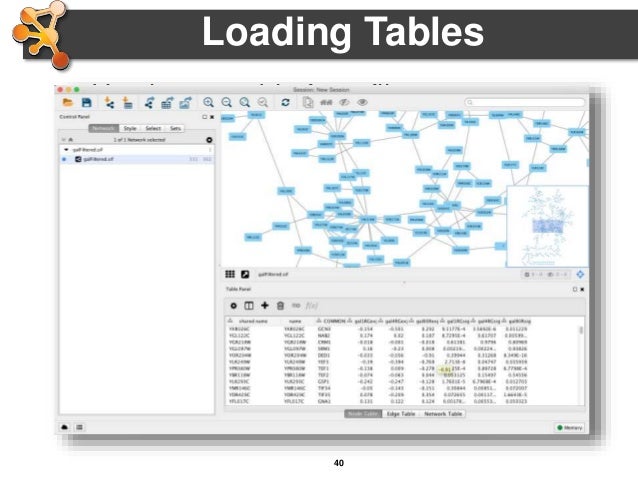
Edit à Preferences à Properties to edit preferences for properties and plugins.All deleted nodes and edges can be restored to the network via Edit à Undo.Options for deleting selected nodes and edges from the current network.Options for creating and destroying views (graphical representations of a network) and networks.Undo and Redo functions which undo and redo edits made in the Attribute Browser, the Network Editor and the Layout.File à Quit closes all windows of Cytoscape and exits the program.File à Export for exporting data and images.File à Import for importing data such as networks and attributes.File à Open for opening a Cytoscape session file.File The File menu contains basic file functionality: Loaded plugins might add tabs to either of these CytoPanels.Ĭytoscape Menus We will briefly run through all the menus available in Cytoscape. The Data Panel starts off with three tabs: Node Attribute Browser, Edge Attribute Browser, and Network Attribute Browser the Network Management panel starts off with four tabs: Network, VizMapper, Editor, and Filters. You can undock any of these panels by clicking on the Float Window control in the upper-‐right corner of the CytoPanel. The Network Management and Data browser panels are dockable tabbed panels known as CytoPanels. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
